Clonal evolution in patients with chronic lymphocytic leukaemia developing resistance to BTK inhibition
Jan A. Burger*^, Dan A. Landau*, Amaro Taylor-Weiner*, Ivana Bozic*, Huidan Zhang*, Kristopher Sarosiek, Lili Wang, Chip Stewart, Jean Fan, Julia Hoellenriegel, Mariela Sivina, Adrian M. Dubuc, Cameron Fraser, Yulong Han, Shuqiang Li, Kenneth J. Livak, Lihua Zou, Youzhong Wan, Sergej Konoplev, Carrie Sougnez, Jennifer R. Brown, Lynne V. Abruzzo, Scott L. Carter, Michael J. Keating, Matthew S. Davids, William G. Wierda, Kristian Cibulskis, Thorsten Zenz, Lillian Werner, Paola Dal Cin, Peter Kharchencko, Donna Neuberg, Hagop Kantarjian, Eric Lander, Stacey Gabriel, Susan O’Brien, Anthony Letai, David A. Weitz, Martin A. Nowak, Gad Getz, Catherine J. Wu^
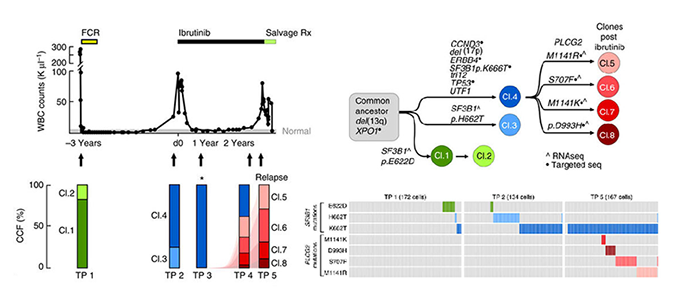
Abstract: Resistance to the Bruton’s tyrosine kinase (BTK) inhibitor ibrutinib has been attributed solely to mutations in BTK and related pathway molecules. Using whole-exome and deep-targeted sequencing, we dissect evolution of ibrutinib resistance in serial samples from five chronic lymphocytic leukaemia patients. In two patients, we detect BTK-C481S mutation or multiple PLCG2 mutations. The other three patients exhibit an expansion of clones harbouring del(8p) with additional driver mutations (EP300, MLL2 and EIF2A), with one patient developing trans-differentiation into CD19-negative histiocytic sarcoma. Using droplet-microfluidic technology and growth kinetic analyses, we demonstrate the presence of ibrutinib-resistant subclones and estimate subclone size before treatment initiation. Haploinsufficiency of TRAIL-R, a consequence of del(8p), results in TRAIL insensitivity, which may contribute to ibrutinib resistance. These findings demonstrate that the ibrutinib therapy favours selection and expansion of rare subclones already present before ibrutinib treatment, and provide insight into the heterogeneity of genetic changes associated with ibrutinib resistance.
Paper: Nature Communications. May 20, 2016. doi:10.1038/ncomms11589
| Pubmed
| PDF
Relevant code: