Transcriptomic Characterization of SF3B1 Mutation Reveals Its Pleiotropic Effects in Chronic Lymphocytic Leukemia
Lili Wang*, Angela N. Brooks*, Jean Fan*, Youzhong Wan*, Rutendo Gambe, Shuqiang Li, Sarah Hergert, Shanye Yin, Samuel S. Freeman, Joshua Z. Levin, Lin Fan, Michael Seiler, Silvia Buonamici, Peter G. Smith, Kevin F. Chau, Carrie L. Cibulskis, Wandi Zhang, Laura Z. Rassenti, Emanuela M. Ghia, Thomas J. Kipps, Stacey Fernandes, Donald B. Bloch, Dylan Kotliar, Dan A. Landau, Sachet A. Shukla, Jon C. Aster, Robin Reed, David S. DeLuca, Jennifer R. Brown, Donna Neuberg, Gad Getz, Kenneth J. Livak, Matthew M. Meyerson, Peter V. Kharchenko, Catherine J. Wu^
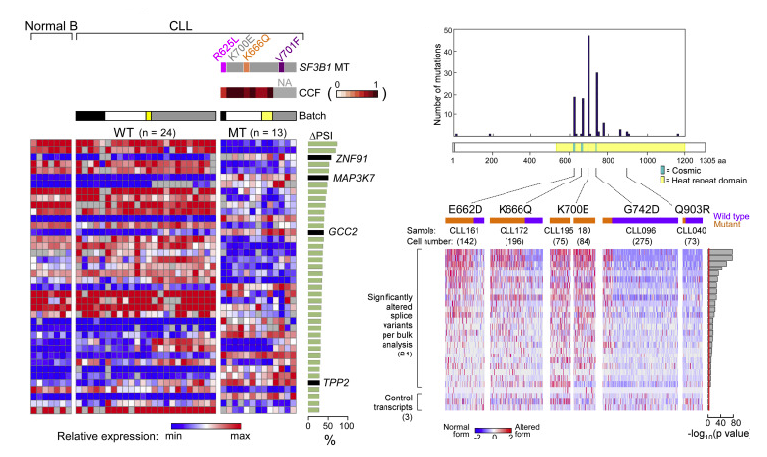
Abstract: Mutations in SF3B1, which encodes a spliceosome component, are associated with poor outcome in chronic lymphocytic leukemia (CLL), but how these contribute to CLL progression remains poorly understood. We undertook a transcriptomic characterization of primary human CLL cells to identify transcripts and pathways affected by SF3B1 mutation. Splicing alterations, identified in the analysis of bulk cells, were confirmed in single SF3B1-mutated CLL cells and also found in cell lines ectopically expressing mutant SF3B1. SF3B1 mutation was found to dysregulate multiple cellular functions including DNA damage response, telomere maintenance, and Notch signaling (mediated through KLF8 upregulation, increased TERC and TERT expression, or altered splicing of DVL2 transcript, respectively). SF3B1 mutation leads to diverse changes in CLL-related pathways.
Paper: Cancer Cell. November 3, 2016. doi.org/10.1016/j.ccell.2016.10.005
| Pubmed
| PDF