Integrated single-cell genetic and transcriptional analysis suggests novel drivers of chronic lymphocytic leukemia
Lili Wang*, Jean Fan*, Joshua M. Francis, George Georghiou, Sarah Hergert, Shuqiang Li, Rutendo Gambe, Chensheng W. Zhou, Chunxiao Yang, Sheng Xiao, Paola Dal Cin, Michaela Bowden, Dylan Kotliar, Sachet A. Shukla, Jennifer R. Brown, Donna Neuberg, Dario R. Alessi, Cheng-Zhong Zhang^, Peter V. Kharchenko, Kenneth J. Livak, Catherine J. Wu^
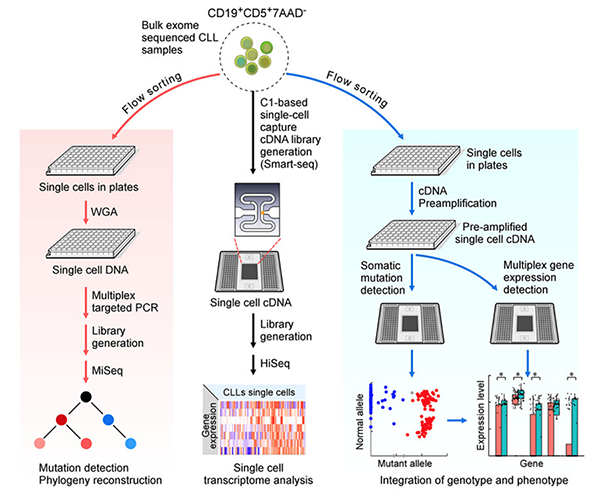
Abstract: Intra-tumoral genetic heterogeneity has been characterized across cancers by genome sequencing of bulk tumors, including chronic lymphocytic leukemia (CLL). In order to more accurately identify subclones, define phylogenetic relationships, and probe genotype–phenotype relationships, we developed methods for targeted mutation detection in DNA and RNA isolated from thousands of single cells from five CLL samples. By clearly resolving phylogenic relationships, we uncovered mutated LCP1 and WNK1 as novel CLL drivers, supported by functional evidence demonstrating their impact on CLL pathways. Integrative analysis of somatic mutations with transcriptional states prompts the idea that convergent evolution generates phenotypically similar cells in distinct genetic branches, thus creating a cohesive expression profile in each CLL sample despite the presence of genetic heterogeneity. Our study highlights the potential for single-cell RNA-based targeted analysis to sensitively determine transcriptional and mutational profiles of individual cancer cells, leading to increased understanding of driving events in malignancy.
Paper: Genome Research. May 22, 2017. doi/10.1101/gr.217331.116
| Pubmed
| PDF