Spatial organization of the transcriptome in individual neurons
Guiping Wang, Cheen-Euong Ang, Jean Fan, Andrew Wang, Jeffrey R. Moffitt, Xiaowei Zhuang^
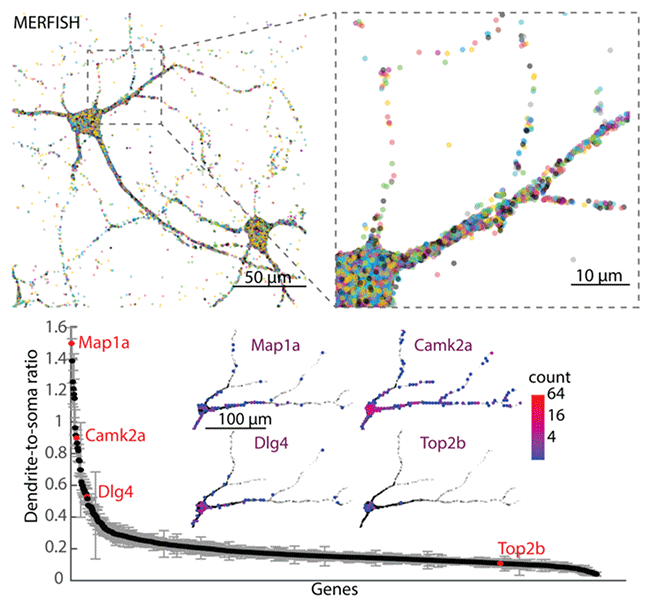
Abstract: Neurons are highly polarized cells with complex neurite morphology. Spatial organization and local translation of RNAs in dendrites and axons play an important role in many neuronal functions. Here we performed super-resolution spatial profiling of RNAs inside individual neurons at the genome scale using multiplexed error-robust fluorescence in situ hybridization (MERFISH), and mapped the spatial organization of up to ~4,200 RNA species (genes) across multiple length scales, ranging from sub-micrometer to millimeters. Our data generated a quantitative intra-neuronal atlas of RNAs with distinct transcriptome compositions in somata, dendrites, and axons, and revealed diverse sub-dendritic distribution patterns of RNAs. Moreover, our spatial analysis identified distinct groups of genes exhibiting specific spatial clustering of transcripts at the sub-micrometer scale that were dependent on protein synthesis and differentially dependent on synaptic activity. Overall, these data provide a rich resource for characterizing the subcellular organization of the transcriptome in neurons with high spatial resolution.
Paper: bioRxiv