Impact of Normalization on tSNE in Kidney Spatial Transcriptomics Data
What happens if I do or not not normalize and/or transform the gene expression data (e.g. log and/or scale) prior to dimensionality reduction?
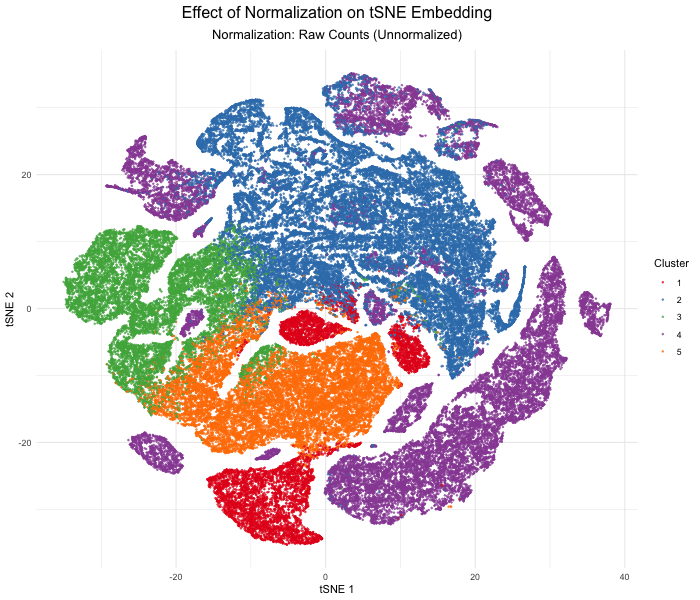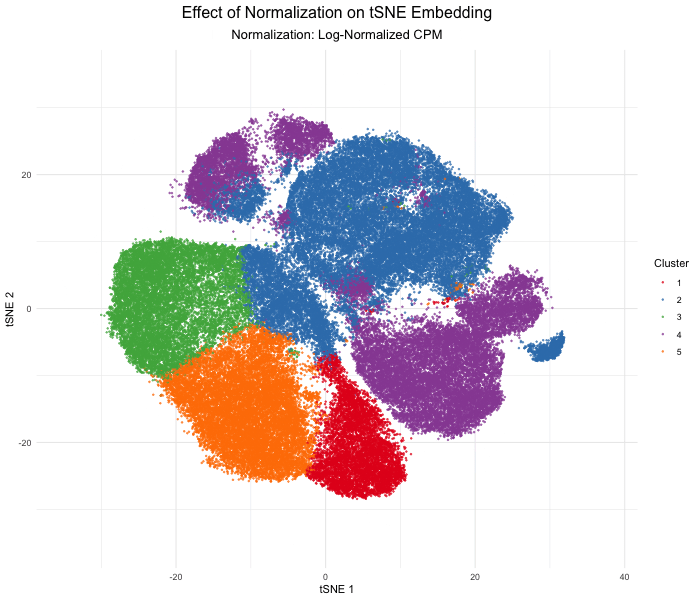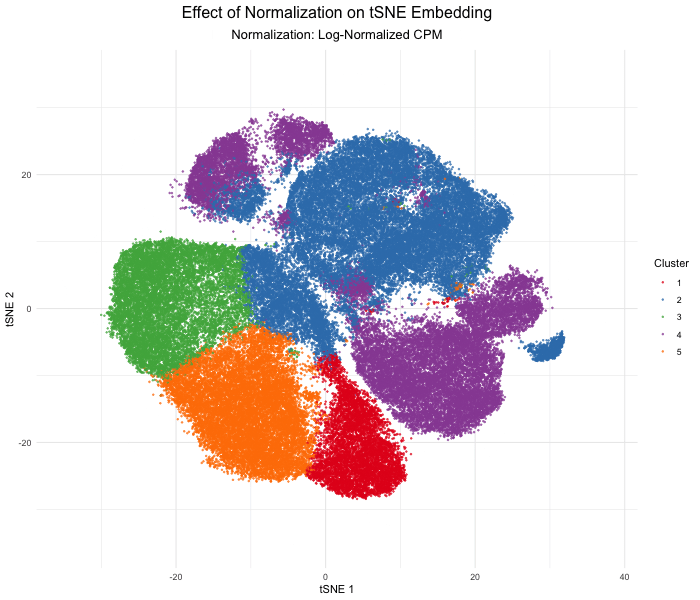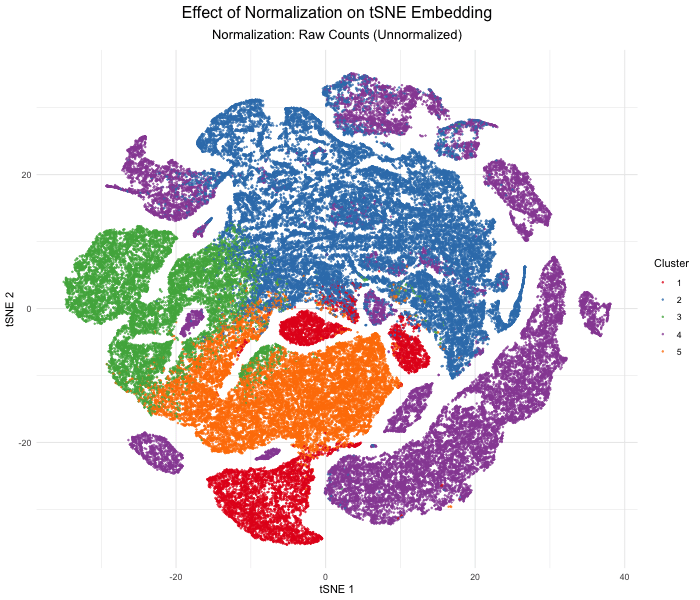
In this analysis, I compared tSNE embeddings of the Xenium single-cell resolution kidney spatial transcriptomics dataset between unnormalized (raw) versus log-normalized counts per million values. The visualization produced above with gganimate explores how normalizing the gene expression data affects dimensionality reduction. Without normalization, the tSNE embedding produced extremely spread out clusters with intermixed clusters, likely due to technical differences rather than true biological differences. In contrast, after log normalization, which divides by the total gene counts, the tSNE embedding produced compact, well-separated clusters that reflect the actual transcriptional differences between cell types. This animated data visualization emphasizes the importance of normalizing gene expression data before dimensionality reduction to avoid technical artifacts from impacting our interpretation of the underlying biology.
Code (paste your code in between the ``` symbols)
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
# Load in Data (Single-cell Resolution Xenium Data) ---------------------------------------------
data <- read.csv('/Users/gracexu/genomic-data-visualization-2026/data/Xenium-IRI-ShamR_matrix.csv')
pos <- data[,c('x', 'y')]
rownames(pos) <- data[,1]
gexp <- data[, 4:ncol(data)]
rownames(gexp) <- data[,1]
gexp[1:5,1:5]
dim(gexp)
# Load in required packages ------------------------------------------------------------------------------------------
library(ggplot2)
library(patchwork)
library(gganimate)
library(gifski)
# Create Conditions: Normalized vs. No Normalization
totgexp <- rowSums(gexp)
# 1. Raw Counts (no normalization)
mat_raw <- gexp
# 2. Log-Normalized Counts Per Million
mat_log <- log10(gexp / totgexp * 1e6 + 1)
# PCA (Linear dimension reduction) & tSNE ------------------------------------------------------------------------------------------
## Write a function to perform PCA + tSNE for each condition
run_tsne <- function(mat, seed = 123) { # include this in the attributes
pcs <- prcomp(mat, center = TRUE, scale = FALSE)
toppcs <- pcs$x[, 1:10]
set.seed(seed)
tsne <- Rtsne::Rtsne(toppcs, dims = 2, perplexity = 30)
tsne$Y
}
emb_raw <- run_tsne(mat_raw)
emb_log <- run_tsne(mat_log)
# k-Means Clustering (base the clustering labels on the normalized data) ------------------------------------------------------------------------------------------------------------------------
toppcs_log <- prcomp(mat_log, center = TRUE, scale = FALSE)$x[, 1:10]
set.seed(123)
km <- kmeans(toppcs_log, centers = 5)
names(km)
cluster <- as.factor(km$cluster)
# Combine into one dataframe ------------------------------------------------------------------------------------------------------------------------
df <- function(emb, condition_label) {
data.frame(
cell = rownames(gexp),
X1 = emb[, 1],
X2 = emb[, 2],
cluster = cluster,
condition = condition_label
)
}
df_combined <- rbind(
df(emb_raw, "Raw Counts (Unnormalized)"),
df(emb_log, "Log-Normalized CPM")
)
df_combined$condition <- factor(df_combined$condition,
levels = c("Raw Counts (Unnormalized)", "Log-Normalized CPM"))
# Create an Animated Plot ------------------------------------------------------------------------------------------------------------------------
p_anim <- ggplot(df_combined, aes(x = X1, y = X2, col = cluster)) +
geom_point(size = 0.4, alpha = 0.6) +
scale_color_brewer(palette = "Set1") +
labs(
title = "Effect of Normalization on tSNE Embedding",
subtitle = "Normalization: {closest_state}",
x = "tSNE 1",
y = "tSNE 2",
color = "Cluster"
) +
theme_minimal() +
theme(
plot.title = element_text(hjust = 0.5, size = 16),
plot.subtitle = element_text(hjust = 0.5, size = 13, face = "bold")
) +
transition_states(
condition,
transition_length = 2,
state_length = 3
) +
ease_aes('cubic-in-out')
# Animate
animate(
p_anim,
nframes = 90,
fps = 15,
width = 700,
height = 600,
renderer = gifski_renderer()
)
# Save plot
anim_save("/Users/gracexu/genomic-data-visualization-2026/homework_submission/HW3_EC_normalization.gif")
# AI Help Prompts:
## I have two tSNE embeddings from different normalization conditions in R. How can I combine them into a single dataframe with a condition label column
## How can I use gganimate to animate between two different tSNE embedding visualizations, where each state represents a different normalization condition (normalized vs. raw data), and also have the subtitle automatically update to show the current normalization state?
