tSNE on varying PC numbers
Description
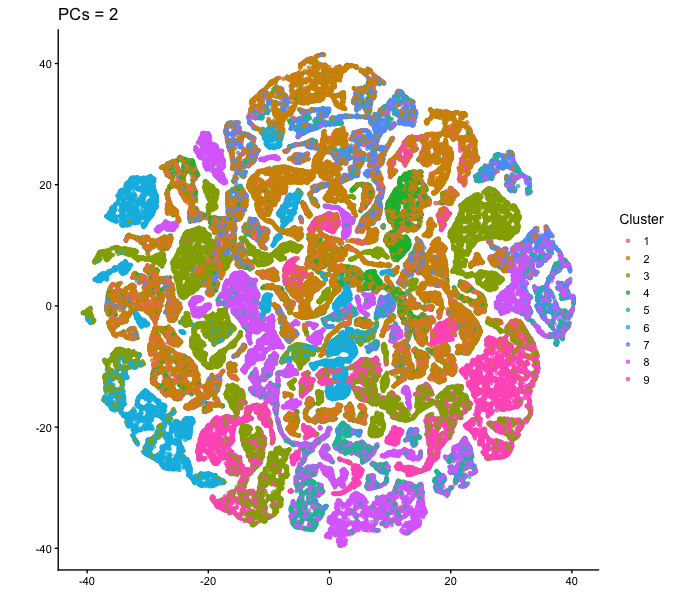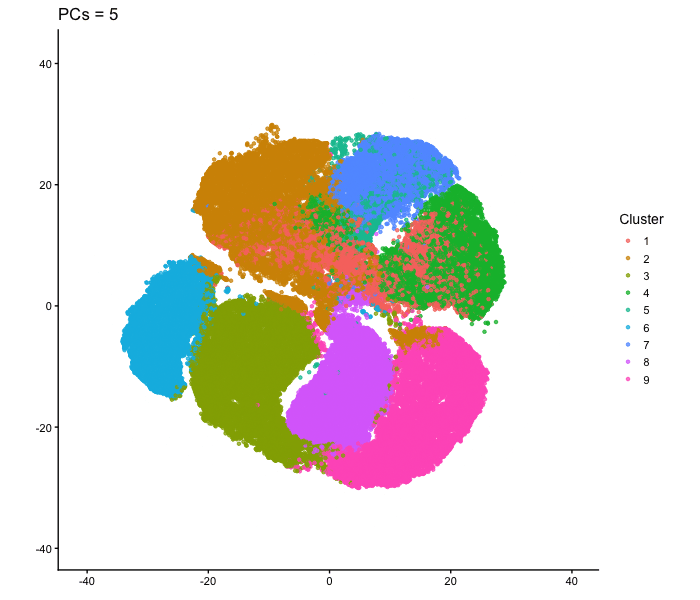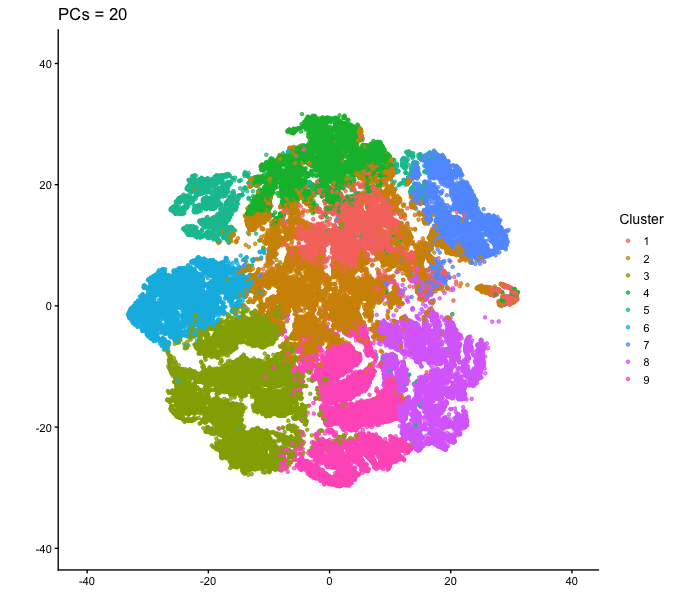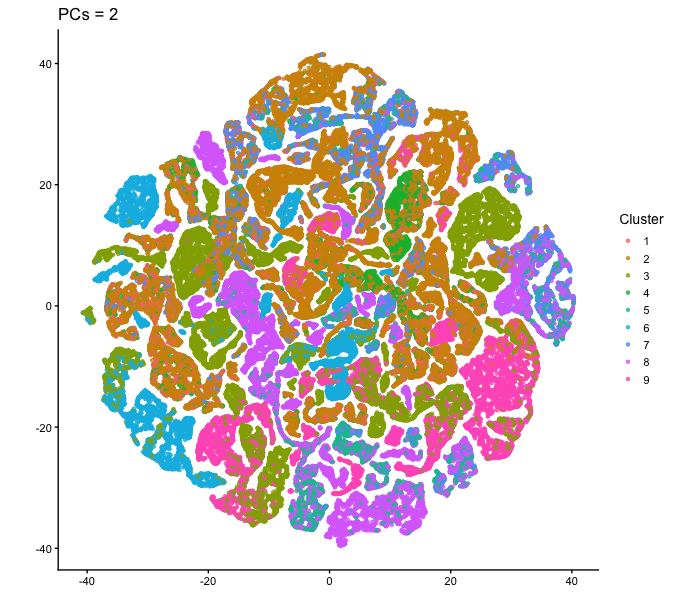
This animation adresses the question: “If I perform non-linear dimensionality reduction on PCs, what happens when I vary how many PCs I use?”
To show how the number of PCs affects nonlinear dimensionality reduction, I first performed PCA on the normalized gene expression data and extracted the PC scores. I then selected different numbers of PCs (2, 3, 5, 10, 15, 20, and 30) and used each as input to tSNE. For each one, I made a 2D tSNE embedding and colored the cells according to k-means cluster assignments (based on the first 10 PCs). I combined these embeddings into an animation to show how the overall structure changes as more PCs are included. When only two PCs are used, the embedding looks relatively messy, with clusters overlapping and without any clear separation. As more PCs are added (around 3-5 PCs), clearer structure starts to appear. By the time 10-15 PCs are included, the clusters form well-defined groups. With more PCs than that (20-30 PCs), the overall layout stays mostly similar, with only small changes in cluster shape and positioning instead of completely reorganizing like before. Overall, this animation shows that the number of PCs used as input to tSNE has an impact on the embedding, especially at lower PC counts. Using too few PCs doesn’t have enough information for important structure, but including more PCs allows meaningful patterns to be shown. After a certain point, adding more PCs produces less change, since most of the relevant signal is already captured in the earlier components.
Code (paste your code in between the ``` symbols)
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
#note: this takes really long to run because of doing the tSNE on the full dataset
library(ggplot2)
library(dplyr)
library(Rtsne)
library(gganimate)
data <- read.csv("~/Desktop/Xenium-IRI-ShamR_matrix.csv.gz")
pos <- data[, c("x", "y")]
rownames(pos) <- data[, 1]
gexp <- data[, 4:ncol(data)]
rownames(gexp) <- data[, 1]
totgexp <- rowSums(gexp)
mat <- log10(gexp / totgexp * 1e6 + 1)
vg <- apply(mat, 2, var)
vargenes <- names(sort(vg, decreasing = TRUE)[1:250])
matsub <- mat[, vargenes, drop = FALSE]
pcs <- prcomp(matsub)
pc_scores <- pcs$x
#K-means based on 10 PCs
set.seed(1)
km <- kmeans(pc_scores[, 1:10, drop = FALSE], centers = 9)
clusters <- factor(km$cluster)
#Compute tSNE
pc_sets <- c(2, 3, 5, 10, 15, 20, 30)
perplexity_val <- 30
tsne_list <- lapply(pc_sets, function(n_pcs) {
set.seed(1)
ts <- Rtsne(
pc_scores[, 1:n_pcs, drop = FALSE],
dims = 2,
perplexity = perplexity_val,
check_duplicates = FALSE,
verbose = FALSE
)
#added this because it takes forever to run so I can see how far it is
message("Made it through PCs = ", n_pcs)
data.frame(
tSNE1 = ts$Y[, 1],
tSNE2 = ts$Y[, 2],
cluster = clusters,
pc_state = factor(paste0("PCs = ", n_pcs), levels = paste0("PCs = ", pc_sets))
)
})
tsne_anim_df <- bind_rows(tsne_list)
#Fix axis limits so the view doesn't jump ----
x_lim <- range(tsne_anim_df$tSNE1, na.rm = TRUE)
y_lim <- range(tsne_anim_df$tSNE2, na.rm = TRUE)
#build animation
anim <- ggplot(tsne_anim_df, aes(tSNE1, tSNE2, color = cluster)) +
geom_point(size = 1.2, alpha = 0.75) +
theme_classic(base_size = 14) +
coord_fixed(xlim = x_lim, ylim = y_lim) +
labs(
title = "{closest_state}",
x = NULL,
y = NULL,
color = "Cluster"
) +
transition_states(
pc_state,
transition_length = 2,
state_length = 3
) +
ease_aes("cubic-in-out")
#render
animate(anim, nframes = 300, fps = 10, width = 700, height = 600, renderer = gifski_renderer())
#save
anim_save("tsne_pc_animation.gif", animate(anim, nframes = 300, fps = 10, width = 700, height = 600, renderer = gifski_renderer()))
Attributions: Code adapted from in-class demo on gganimate. AI used for debugging and for function to loop through PC values.
