Characterizing spatial gene expression heterogeneity in spatially resolved single-cell transcriptomics data with nonuniform cellular densities
Brendan F Miller, Dhananjay Bambah-Mukku, Catherine Dulac, Xiaowei Zhuang, Jean Fan^
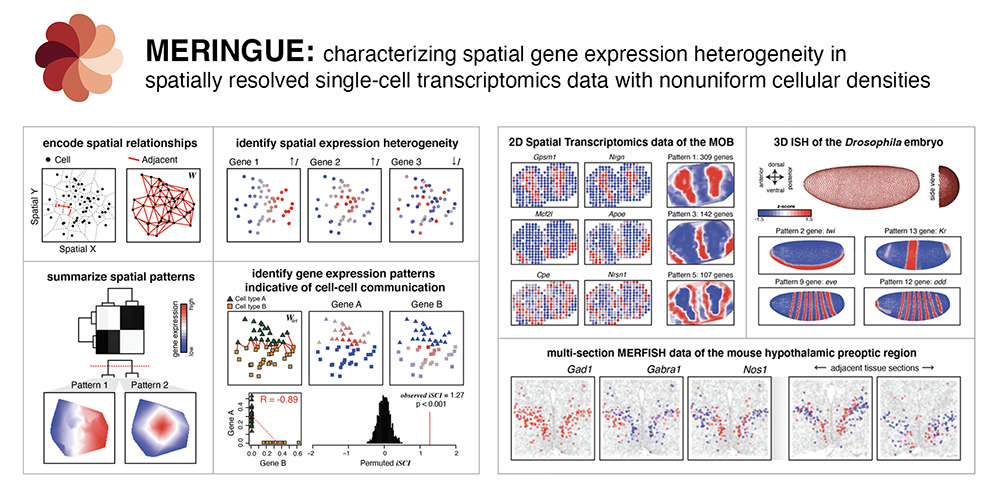
Abstract: Recent technological advances have enabled spatially resolved measurements of expression profiles for hundreds to thousands of genes in fixed tissues at single-cell resolution. However, scalable computational analysis methods able to take into consideration the inherent 3D spatial organization of cell types and nonuniform cellular densities within tissues are still lacking. To address this, we developed MERINGUE, a computational framework based on spatial auto-correlation and cross-correlation analysis to identify genes with spatially heterogeneous expression patterns, infer putative cell-cell communication, and perform spatially informed cell clustering in 2D and 3D in a density-agnostic manner using spatially resolved transcriptomics data. We applied MERINGUE to a variety of spatially resolved transcriptomics datasets including multiplexed error-robust fluorescence in situ hybridization (MERFISH), spatial transcriptomics, Slide-Seq, and aligned in situ hybridization (ISH) data. We anticipate that such statistical analysis of spatially resolved transcriptomics data will facilitate our understanding of the interplay between cell state and spatial organization in tissue development and disease.
Paper: Genome Research. Advance May 25, 2021, doi:10.1101/gr.271288.120
| Pubmed
| PDF
Relevant code: